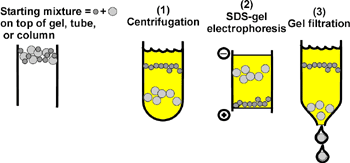
Methods based on size for separation of large molecules
1. Centrifugation
When you put a particle in a centrifugal field, it is acted upon by the centrifugal force, which is proportional to the molecular weight (M), to the square of the speed (angular velocity, rpm) of the rotor (w2) and to the distance of the solution from the center of rotation (r): Cent. Force ~ Mw2r. Acting in the opposite direction to particle motion is friction. This frictional force is proportional to the velocity of the particle (V) and to a coefficient of friction (f) that depends on the shape of the particle: Friction force ~ fV. The particle will accelerate until a velocity is achieved such that these two opposing forces are equal, after which the particle will continue to sediment, but at a constant velocity. Setting the 2 forces equal to each other we have: Mw2r=fV, and so:
V ~ Mw2r/f.
(NOTE that this equation was printed incorrectly with w2r in the denominator on the hard copy of this handout).
In general, this means that velocity increases as mass (molecular weight) increase, and velocity decreases as the cross-sectional size (diameter) increases (f increases with cross-sectional size).
V depends on the particular machine and on the speed of rotation. So you can compare results irregardless of machine and speed, you calculate the Svedberg constant or S value instead of V. S = V/w2r. Macromolecules usually have S values betw een 1 and 100 X 10-13 sec. A value of 1 X 10-13 sec is called one Svedberg unit or S. Many molecules are known by their S values, for example 23 S ribosomal RNA. Its S value was known before its function or molecular weight, and so has become its name.
Molecules of different S values are usually separated by layering them on top of a dense solution (such as sucrose) and then running the centrifuge to force the molecules through the solution. The molecules of larger S value go farther because of eithe r larger molecular weight or less frictional drag or both. You stop the centrifuge when the different molecules have traveled different distances down the tube and then remove the solution by puncturing the bottom of the tube or pumping the solution off t he top. The invention of the ultracentrifuge (a super fast centrifuge) by Svedberg was what originally made the purification of macromolecules possible. Today there are many other methods also used, as listed below.
2. Gel Electrophoresis plus SDS
When proteins and other macromolecules are treated with SDS, a strong detergent, they are denatured. They also become negatively charged because of the charge on the detergent, and the amount of detergent bound is so large that any differences in n ative charge are swamped. The bigger the macromolecule, the more SDS is bound, so that all macromolecules treated with SDS have the same ratio of charge to mass. (The amount bound is proportional to mass.) So for molecules treated with SDS, the pull per u nit mass in an electric field is the same, and all molecules should have the same velocity IF there is no frictional drag. But electrophoresis with SDS is always done in a gel, so that the molecules must be pulled though the pores. And the ease of moving through the pores depends on the diameter of the molecules. The bigger molecules are retarded or keep getting stuck and don't move as fast. Since the molecules are all denatured into random coils, the diameter strictly depends on the length or molecular w eight. The bigger the molecular weight, the longer the coil and the slower the molecule goes. So, electrophoresis + SDS separates on the basis of molecular weight, not on the basis of native charge. Important note: Proteins of the same length usually cann ot be separated by gel electrophoresis + SDS. The differences in molecular weight caused by differences in the R groups are not enough to allow a separation. If two proteins migrate the same +SDS they are assumed to be of approximately the same molecular weight because they are about the same length (contain the same # of amino acids).
3. Gel filtration or molecular sieve chromatography
You can make a column packed with beads that have holes in them. The beads are made of a gel, and the holes can be any size you want, within limits. There are many kinds of beads commercially available, but the most famous is called Sephadex. You f ill up a column with the beads, and then fill the spaces around the beads with a buffer. Then you put your mixture of molecules on the top and wash it through the column. The smaller molecules that can fit through the holes in the beads take longer to tra vel the length of the column. The larger molecules that can not enter the beads at all move only through the spaces between the beads and come quickly out the other end. So smaller molecules take LONGER to come out than bigger ones. Note this is the oppos ite of gel electrophoresis. The results are different here because you have a two phase system - the big molecules are totally excluded from the small holes in the beads, not forced to travel through tight spaces as in gel electrophoresis. This method sep arates on the basis of size or diameter. If diameter is proportional to molecular weight, as it often is, this method can be used to estimate molecular weights or to separate on the basis of molecular weight. The buffer used here can be one that is consis tent with the maintenance of tertiary and quaternary structure, so this method is useful for determining the MW of native proteins.
